Welcome to IMDA’s documentation!
The ImmunoDataAnalyzer (IMDA) pipeline provides various methods for analysing immunological next-generation sequencing (NGS) data. IMDA allows to process data from immunoglobulins (IG) and T cell receptors (TCR). The pipeline is especially build for first interpretations and quality control of TCR or IG data. Several aspects based on CDR3 and V(D)J genes are calculated, evaluated and visualizations are provided in a compact format.
The software is developed by researchers of the Bioinformatics Research Group at FH Upper Austria, Hagenberg Campus and researchers of the Division of Nephrology and Dialysis, Department of Medicine III, Medical University of Vienna.
Getting Started
How to use IMDA
Tutorial
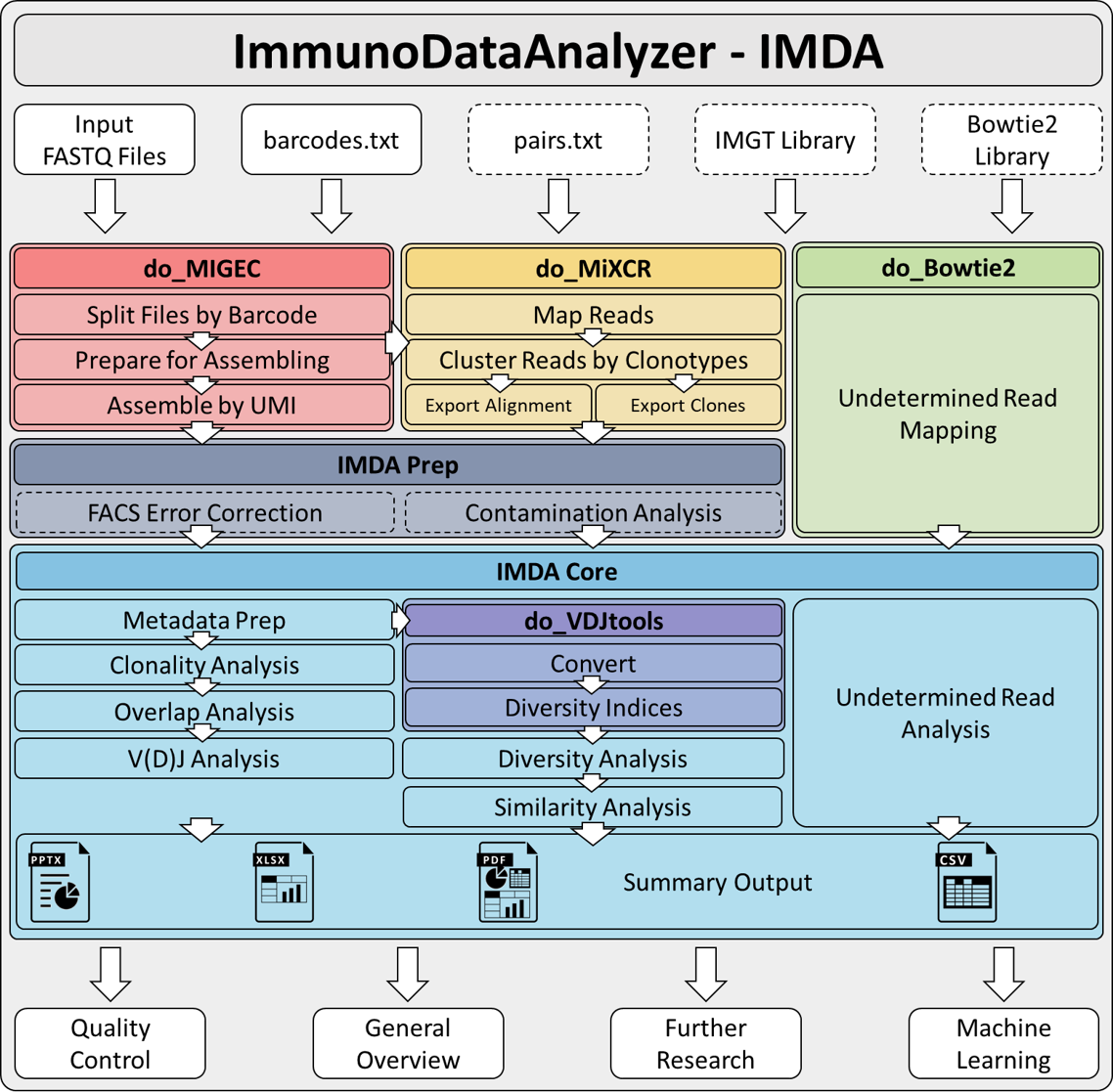
IMDA Pipeline
Additional Information