Data Science


In the research area Data Science, we focus on the development and application of data-driven methods for solving biological, medical, and technical problems. In close cooperation with our partners from research and industry, we develop algorithms for preprocessing and analyzing data using statistical methods and machine learning, as well as visualizing and interpreting the results. For example, within the project LeiVMed Online we develop a platform for benchmarking and visualizing data of treatments of patients in hospitals and developing prediction models for the outcome of treatments.
In many of our projects, we see the need for customized data processing pipelines due to the heterogeneity and complexity of data structures in real world processes and systems. We use applied statistics as well as numerous machine learning approaches, including black box methods (deep neural nets, random forests, gradient boosted trees, etc.) and white box methods (symbolic regression by genetic programming). We build our knowledge discovery pipelines on a variety of different frameworks, especially python scikit-learn, tensorflow, pytorch, MATLAB, and HeuristicLab.
Selected Publications
| 2021 | Stefan Anlauf, Andreas Haghofer, Karl Dirnberger, Stephan M. Winkler: | Data-Based Prediction of Microbial Contamination in Herbs and Identification of Optimal Harvest Parameters | International Journal of Food Engineering (IJFE), DeGruyter | View Article |
| 2019 | Renate Haselgrübler, Peter Lanzerstorfer, Clemens Röhrl, Flora Stübl, Jonas Schurr, Bettina Schwarzinger, Clemens Schwarzinger, Mario Brameshuber, Stefan Wieser, Stephan M. Winkler, and Julian Weghuber: | Hypolipidemic effects of herbal extracts by reduction of adipocyte differentiation, intracellular neutral lipid content, lipolysis, fatty acid exchange and lipid droplet motility | Scientific Reports | View Article |
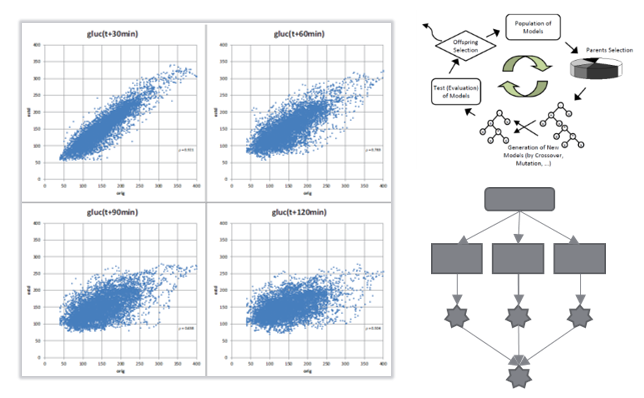
| 2018 | Stephan M. Winkler: | Evolutionary Computation and Symbolic Regression in Scientific Modeling | , Johannes Kepler University Linz, Computer Science, Habilitation Thesis | |
| 2016 | Daniela Borgmann, Sandra Mayr, Helene Polin, Susanne Schaller, Viktoria Dorfer, Christian Gabriel, Stephan Winkler, and Jaroslaw Jacak: | Single Molecule Florescence Microscopy and Machine Learning for Rhesus D Antigen Classification | Scientific Reports 6 | View Article |
Selected Projects
Interoperability Smile
Health organizations around the world spend a lot of time and money calculating quality reports manually. In 2020, Health Level 7 (HL7) introduced the Clinical Quality Language (CQL) which allows for expression of clinical knowledge that can be used in a variety of ways, including cohort definition, clinical decision support, public health reporting and quality measurement.
CQL is meant to be readable by non-technical audience and has the potential to automatically generate quality reports that are more transparent, less ambiguous and ultimately cheaper compared to manually calculated reports.
In collaboration with our Canadian partner Smile CDR Inc., we are working on a proof of concept for using CQL to automatically generate measures on the immunization status of a population. For that, we are using the open-source library HAPI FHIR and sample data from a FHIR-based clinical data repository.
Links:
CQL, SmileCDR, HAPI FHIR, FHIR
Head
Prof.(FH) DI Dr. Stephan Winkler
Researchers
Anna Lin MSc.
Anja Schwab
Duration
03/2022 - present
Research Areas
DataScience
Project Partner
Smile CDR Inc.