Welcome to IMPI’s documentation!¶
Our Interface for Point Mutation Identification (IMPI) is a stand-alone software and implements algoirthms which allow for unique molecular identifier (UMI) tagged next-generation sequencing (NGS) data processing and the analysis of minor and variant allele frequencies (MAF and VAF), respectively.
The software is developed by researchers of the Bioinformatics Research Group at FH Upper Austria, Hagenberg Campus and researchers of the Laboratory for Molecular Genetic Diagnostics at the Ordensklinikum Linz GmbH, Barmherzige Schwestern.
Getting Started
How to use IMPI
Tutorial
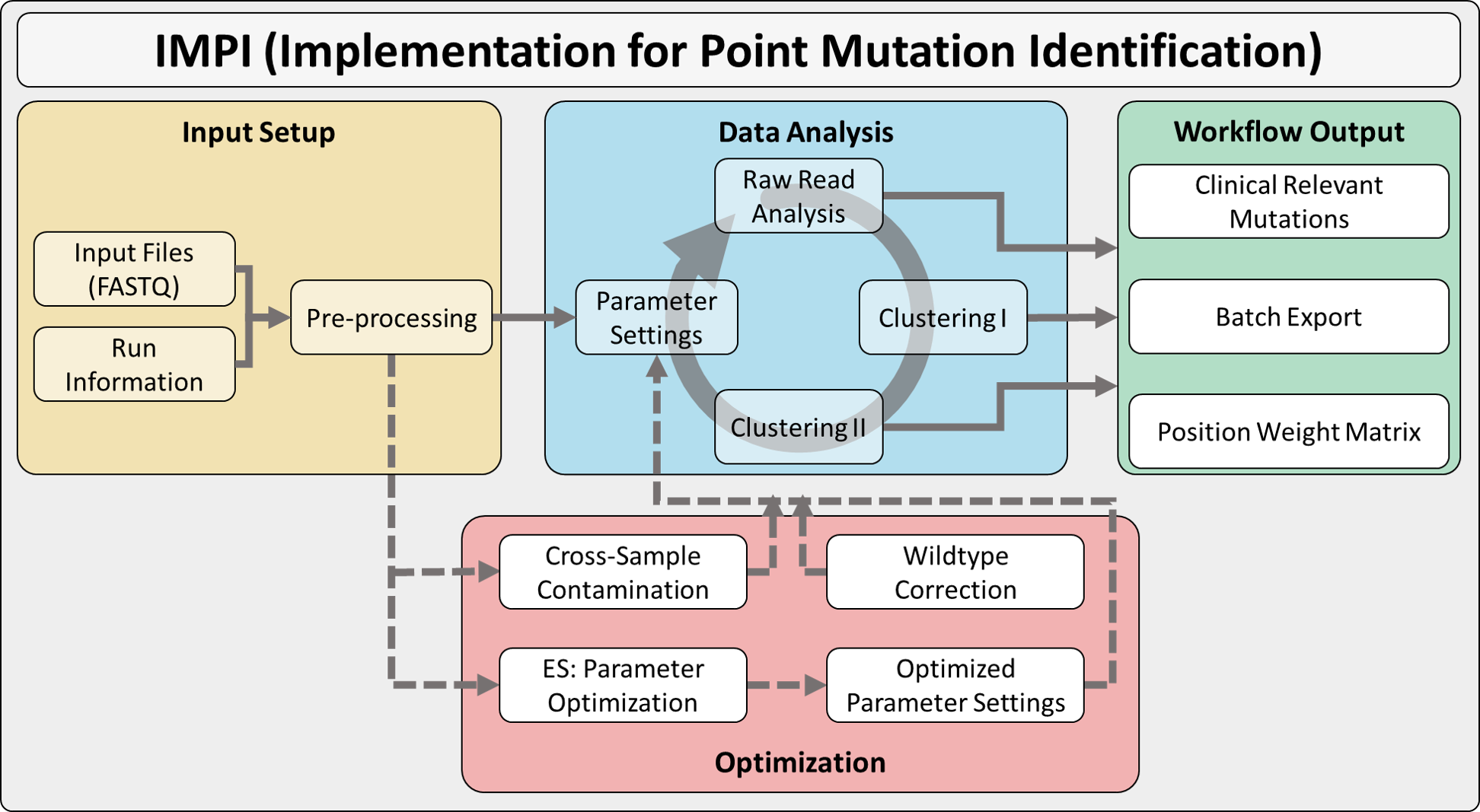
IMPI Workflow¶
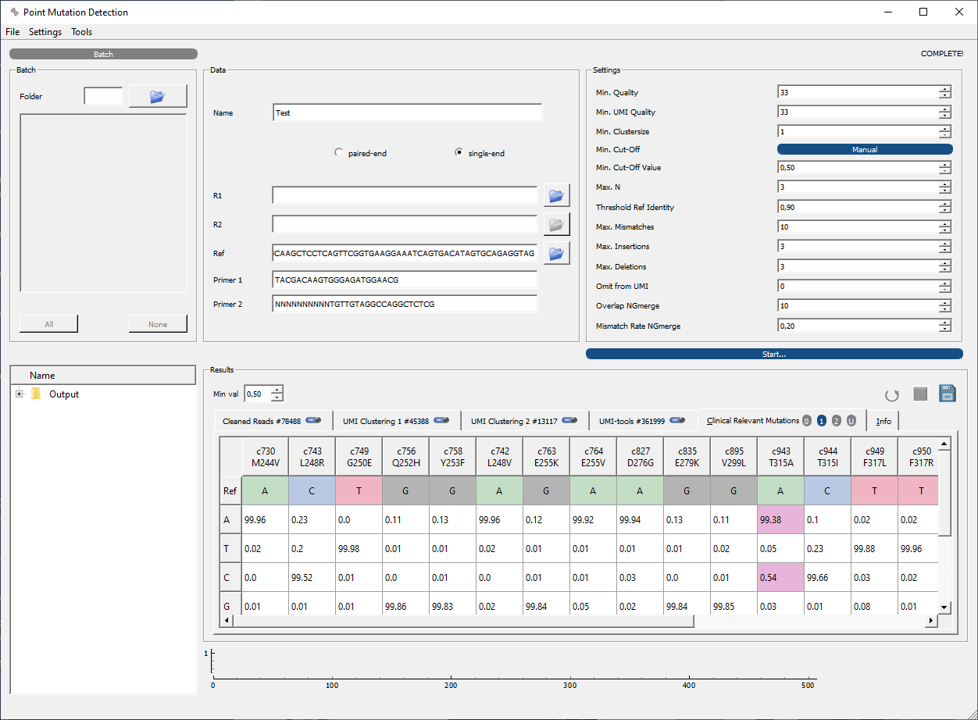
Interface for Point Mutation Identification (IMPI)¶
Additional Information